Home to the Chemical Reaction Database
The chemical reaction database (CRD) is a collection of chemical reactions drawn from the scientific literature and patent literature. Work in progress, for now emphasis on organic reactions
Includes search options for catalysts and ligands, all data normalized with calculated ratio's for each reaction component.
The current database size is over 947 thousand reaction records, over 1.1 million compounds and 300 reaction types (with 528K reactions attributed). The virtual stockroom has 1338 common and less common reagents and 83 solvents.
Reaction SMILES dataset now available on Figshare, see 10.6084/m9.figshare.22491730.v1
2024: added dataset of USPTO 2023 only (137K entries). Including reagents and solvents. 10.6084/m9.figshare.22491730.v1
Papers are considered for inclusion if supplemental info contains individual reaction steps with the systematic name for each compound (Opsin validated) for all reactants and products and clear identifiable reagents. The estimated percentage of papers meeting these requirements in 2022 is smaller than 1%.
New datasets
- (±)-Salvinorin A formal synthesis, H. Tran et al. 2023 reaction data | DOI
- (−)-Cotylenol, S.I. Ting et al. 2023 reaction data | DOI
- Salinosporamide D, S. Park et al. 2023 reaction data | DOI
- 10-Deoxymethynolide, K.J. Chambers et al. 2023 reaction data | DOI
- (+)-Euphorikanin A, M.J. Classen et al. 2023 reaction data | DOI
- Isoxeniolide A, L. Betschart and K-H. Altmann 2023 reaction data | DOI
- pancratistatin framework, D. Azubuike et al. 2024 reaction data | DOI
- sealutomicin C, S. Astle et al. 2024 reaction data | DOI
- Aanacine, G. Han et al. 2024 reaction data | DOI
- Lobatamide C, S. Yasui et al. 2024 reaction data | DOI
 Example organic reaction
Example organic reaction
The blog
Recently in the blog: The year in review and alkyne protiodesilylation by the numbers
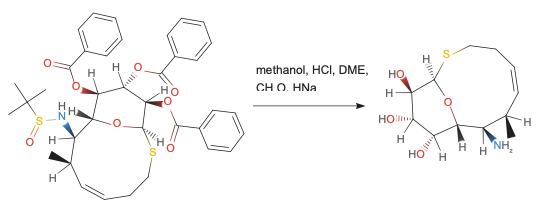 Example organic reaction
Example organic reaction
Organic reactions by year
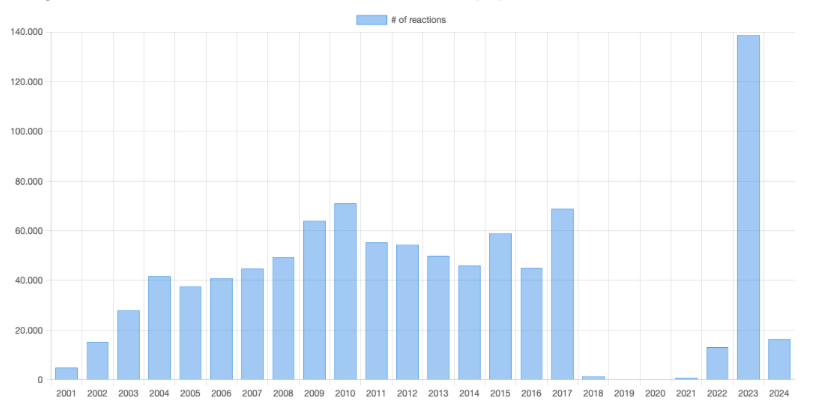 Organic reactions in the database by year
Organic reactions in the database by year
Main datasets
Currently the database contains 4 main datasets. The first is the USPTO dataset 2001-2016 as compiled by Daniel Lowe with data enhancing. The second dataset is also mined from USPTO but with custom programming and with the aid of Oscar4 software or ChatGPT-3.5 and the Opsin service. Currect backlog of years 2018 to 2022. The third dataset is derived from the academic literature (anything with a DOI), progressing at a snail pace (is manual labour). Occasional use of Decimer and Clipboard-To-SMILES Converter. The CJHIF dataset (academic literature) is also included, a total amount of 3.2 million records but only a fraction included thus far. Additional SMILES to IUPAC conversion by STOUT. Reaction images by SmilesDrawer. Reaction types calculated with RDKit.